Editor's note: This story is being highlighted in ASU Now's year in review. Read more top stories from 2018 here.
In a major advancement in nanomedicine, Arizona State University scientists, in collaboration with researchers from the National Center for Nanoscience and Technology (NCNST) of the Chinese Academy of Sciences, have successfully programmed nanorobots to shrink tumors by cutting off their blood supply.
“We have developed the first fully autonomous, DNA robotic system for a very precise drug design and targeted cancer therapy,” said Hao Yan, director of the ASU Biodesign Institute’s Center for Molecular Design and Biomimetics and the Milton Glick Professor in the School of Molecular Sciences.
“Moreover, this technology is a strategy that can be used for many types of cancer, since all solid tumor-feeding blood vessels are essentially the same,” Yan said.
The successful demonstration of the technology, the first-of-its-kind study in mammals utilizing breast-cancer, melanoma, ovarian and lung-cancer mouse models, was published in the journal Nature Biotechnology.
Seek and destroy
Yan is an expert in the field of DNA origami, which in the past two decades has developed atomic-scale manufacturing to build more and more complex structures.
The bricks to build their structures come from DNA, which can self-fold into all sorts of shapes and sizes — all at a scale 1,000 times smaller than the width of a human hair — in the hopes of one day revolutionizing computing, electronics and medicine.
That one day may be coming a bit faster than anticipated.
RELATED: 'DNA origami' is the shape of things to come for nanotechnology
Nanomedicine is a new branch of medicine that seeks to combine the promise of nanotechnology to open up entirely new avenues for treatments, such as making minuscule, molecule-sized nanoparticles to diagnose and treat difficult diseases, especially cancer.
Until now, the challenge of advancing nanomedicine has been difficult because scientists wanted to design, build and carefully control nanorobots to actively seek and destroy cancerous tumors — while not harming any healthy cells.
The international team of researchers overcame this problem by using a seemingly simple strategy to very selectively seek and starve out a tumor.
This work was initiated about five years ago. The NCNST researchers first wanted to specifically cut off tumor blood supply by inducing blood coagulation with high therapeutic efficacy and safety profiles in multiple solid tumors using DNA-based nanocarriers. Yan’s expertise has upgraded the nanomedicine design to be a fully programmable robotic system, able to perform its mission entirely on its own.
“These nanorobots can be programmed to transport molecular payloads and cause on-site tumor blood-supply blockages, which can lead to tissue death and shrink the tumor,” said Baoquan Ding, a professor at the NCNST in Beijing.
Video animation by Jason Drees, Arizona State University
Nanobots to the rescue
To perform their study, the scientists took advantage of a well-known mouse tumor model, where human cancer cells are injected into a mouse to induce aggressive tumor growth.
Once the tumor was growing, the nanorobots were deployed to come to the rescue.
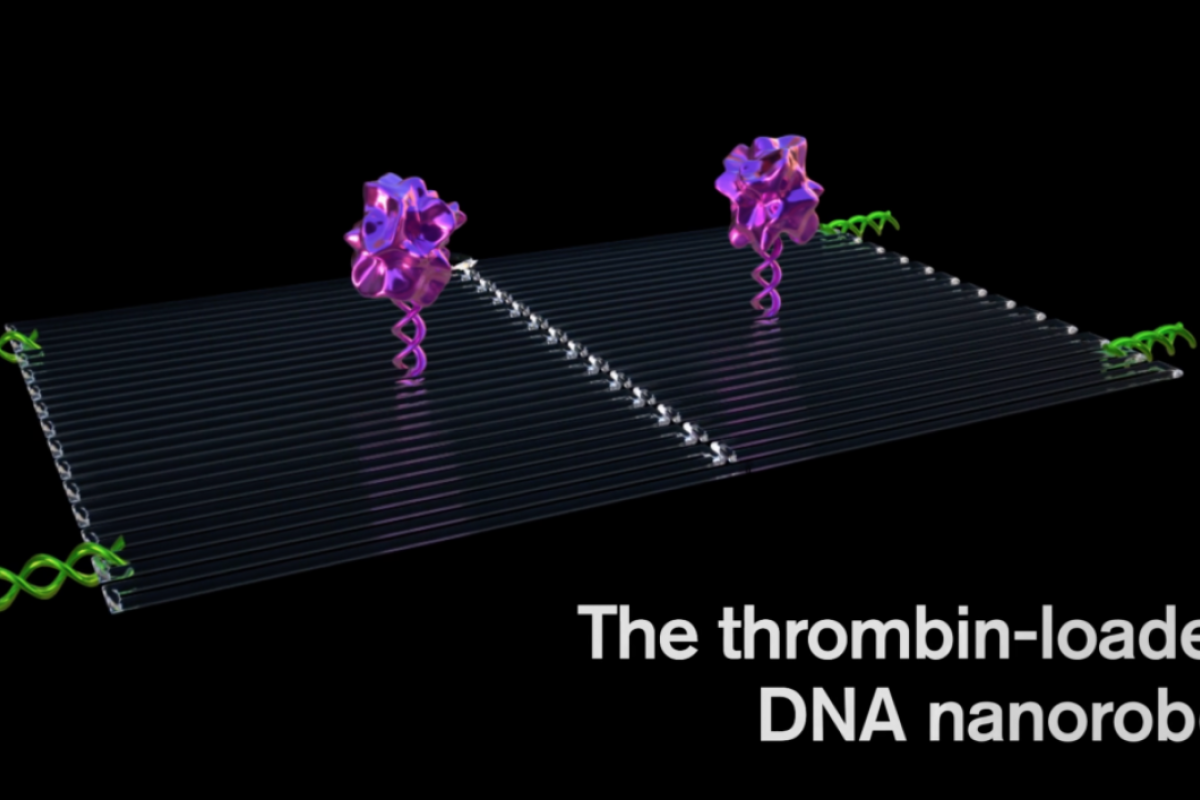
Each nanorobot is made from a flat, rectangular DNA origami sheet, 90 nanometers by 60 nanometers in size. A key blood-clotting enzyme, called thrombin, is attached to the surface.
Thrombin can block tumor blood flow by clotting the blood within the vessels that feed tumor growth, causing a sort of tumor mini heart attack and leading to tumor tissue death.
First, an average of four thrombin molecules was attached to a flat DNA scaffold. Next, the flat sheet was folded in on itself like a sheet of paper into a circle to make a hollow tube.
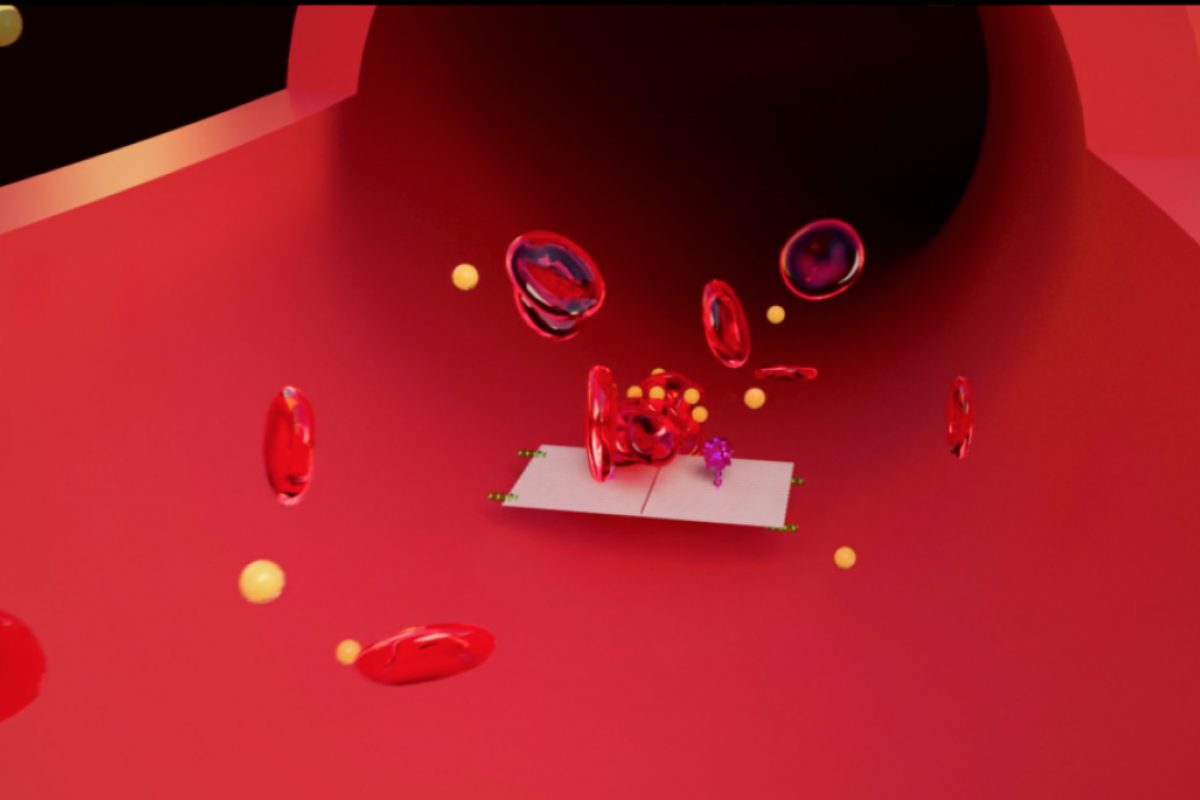
They were injected with an IV into a mouse, then traveled through the bloodstream, homing in on the tumors.
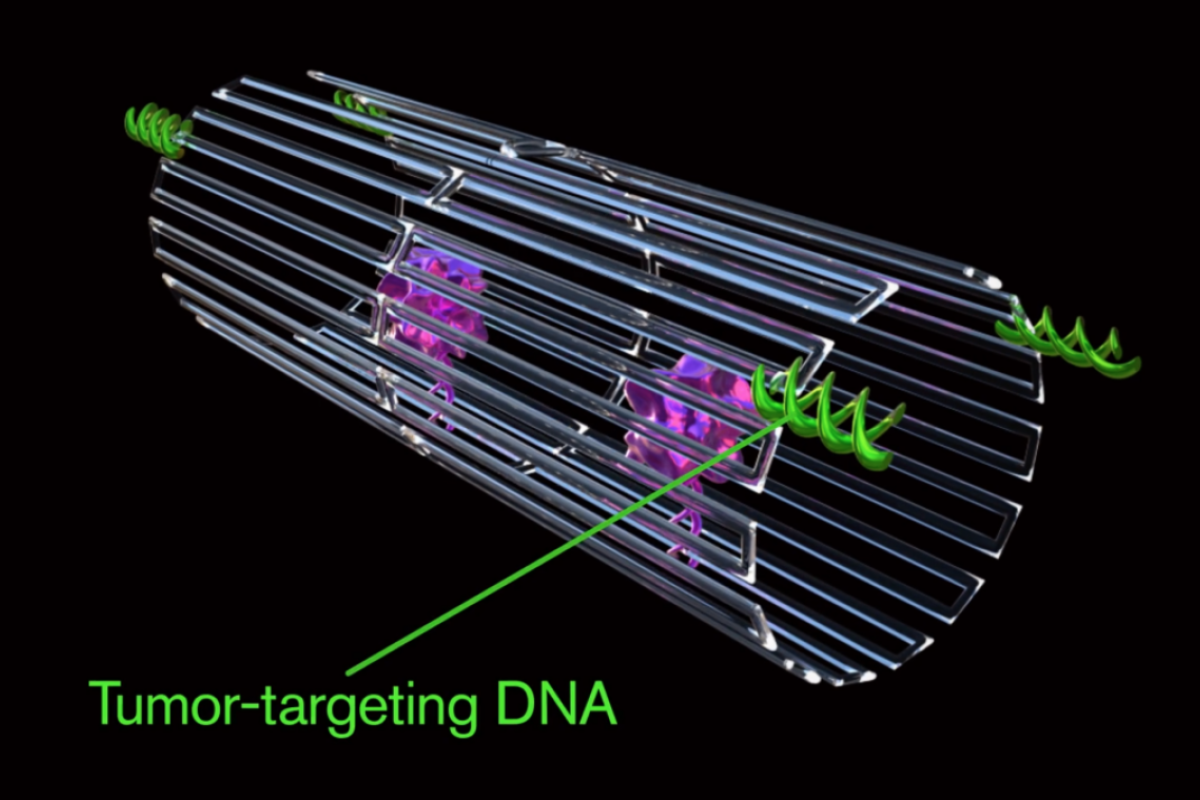
The key to programming a nanorobot that attacks only a cancer cell was to include a special payload on its surface, called a DNA aptamer. The DNA aptamer could specifically target a protein, called nucleolin, that is made in high amounts only on the surface of tumor endothelial cells — and not found on the surface of healthy cells.
Once bound to the tumor blood vessel surface, the nanorobot was programmed, like the notorious Trojan horse, to deliver its unsuspecting drug cargo into the very heart of the tumor, exposing the thrombin.
The nanorobots worked fast, congregating in large numbers to quickly surround the tumor just hours after injection.
Safe and sound design
First and foremost, the team showed that the nanorobots were safe and effective in shrinking tumors.
“The nanorobot proved to be safe and immunologically inert for use in normal mice and, also in Bama miniature pigs, showing no detectable changes in normal blood coagulation or cell morphology,” said Yuliang Zhao, also a professor at NCNST and lead scientist of the international collaborative team.
Most importantly, there was no evidence of the nanorobots spreading into the brain where they could cause unwanted side effects, such as a stroke.
“The nanorobots are decidedly safe in the normal tissues of mice and large animals,” said Guangjun Nie, another professor at the NCNST and a key member of the collaborative team.
The treatment blocked tumor blood supply and generated tumor tissue damage within 24 hours while having no effect on healthy tissues. After attacking tumors, most of the nanorobots were cleared and degraded from the body after 24 hours.
By two days, there was evidence of advanced thrombosisLocal coagulation or clotting of the blood., and at three days, thrombi in all tumor vessels were observed.
The key is to trigger thrombin only when it is inside tumor blood vessels. Also, in the melanoma mouse model, three out of eight mice receiving the nanorobot therapy showed complete regression of the tumors. The median survival time more than doubled, extending from 20.5 to 45 days.
They also tried their system in a test of a primary mouse lung-cancer model, which mimics the human clinical course of lung-cancer patients. They showed shrinkage of tumor tissues after a two-week treatment.
Science of the very small goes big
For Yan, the important study milestone represents the end of the beginning for nanomedicine.
“The thrombin-delivery DNA nanorobot constitutes a major advance in the application of DNA nanotechnology for cancer therapy,” Yan said. “In a melanoma mouse model, the nanorobot not only affected the primary tumor but also prevented the formation of metastasis, showing promising therapeutic potential.”
Yan and his collaborators are now actively pursuing clinical partners to further develop this technology.
“I think we are much closer to real, practical medical applications of the technology,” Yan said. “Combinations of different rationally designed nanorobots carrying various agents may help to accomplish the ultimate goal of cancer research: the eradication of solid tumors and vascularized metastases. Furthermore, the current strategy may be developed as a drug-delivery platform for the treatment of other diseases by modification of the geometry of the nanostructures, the targeting groups and the loaded cargoes.”
This work was supported by grants from National Basic Research Plan of China (MoST Program 2016YFA0201601), the National Natural Science Foundation of China (31730032, 21222311, 21573051, 91127021, the National Distinguished Young Scientists program 31325010), Innovation Research Group of National Natural Science Foundation (11621505, 21721002), Beijing Municipal Science & Technology Commission (Z161100000116035, Z161100000116036), CAS Interdisciplinary Innovation Team, K. C. Wong Education Foundation and US National Institutes of Health Director’s Transformative Research Award (R01GM104960-01).
Top image: Cartoon graphic by Baoquan Ding and Hao Yan
More Science and technology

ASU-led space telescope is ready to fly
The Star Planet Activity Research CubeSat, or SPARCS, a small space telescope that will monitor the flares and sunspot activity of low-mass stars, has now passed its pre-shipment review by NASA.…

ASU at the heart of the state's revitalized microelectronics industry
A stronger local economy, more reliable technology, and a future where our computers and devices do the impossible: that’s the transformation ASU is driving through its microelectronics research…

Breakthrough copper alloy achieves unprecedented high-temperature performance
A team of researchers from Arizona State University, the U.S. Army Research Laboratory, Lehigh University and Louisiana State University has developed a groundbreaking high-temperature copper alloy…