With hospitals more often reaching for antibiotics of last resort to fight infections and recent Ebola and Zika outbreaks crossing borders like never before, the worldwide scientific community has been challenged with developing new antimicrobials to safeguard the population.
The research arm of the U.S. Department of Defense — the Defense Advanced Research Program Agency, or DARPA — is known for taking on outsize challenges such as this. And so, it put out a call for researchers to figure out a process for making at least 1,000 doses to potentially any pathogen within a week of its discovery.
An Arizona State University team was one of the few that rose to the challenge.
“As far as we know, we were the only team to figure out how to do this for any pathogen — virus or bacterium,” said research leader Stephen Albert Johnston, who directs the ASU Biodesign Institute’s Center for Innovations in Medicine and is a professor in the School of Life Sciences. “While the system is designed to create antimicrobials in an extreme emergency — which we hope is never needed — the basic elements can be applied to improve conventional approaches to making anti-infectives.
“My research center thrives on taking on projects most think cannot be done. This challenge was too good not to respond.”
Synthetic sentinels
Antibodies are large, Y-shaped proteins produced by the human immune system to ward off foreign invaders. Our bodies mount this defense rapidly, especially if they have seen the invader before, producing the needed antibody within days of infection.

ASU scientists have recently met a challenge of developing a new class of antimicrobials, called synbodies, to safeguard the population against infectious threats.
But to make antibodies in the lab specific for just one invader can take months, and it can be an expensive proposition.
Johnston wanted to mimic nature’s approach while dramatically reducing the antimicrobial discovery and production time.
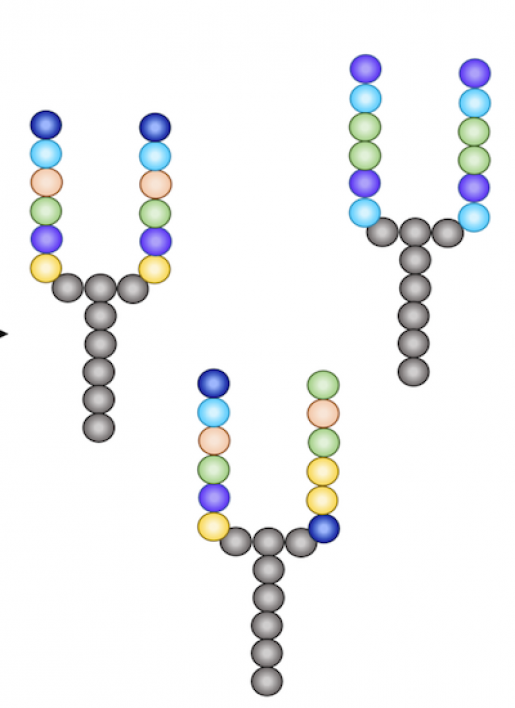
For the past decade, Johnston’s team has been a pioneer in developing lab-made versions that focus on just the business end of antibodies, critical pathogen recognition elements called synthetic antibodies, or synbodies.
Synbodies are made from two short protein fragments, called peptides, which are joined together to form a small, antibody-like compound still large enough to do its job.
Chipping away
But even the process of making synbodies typically takes several months.
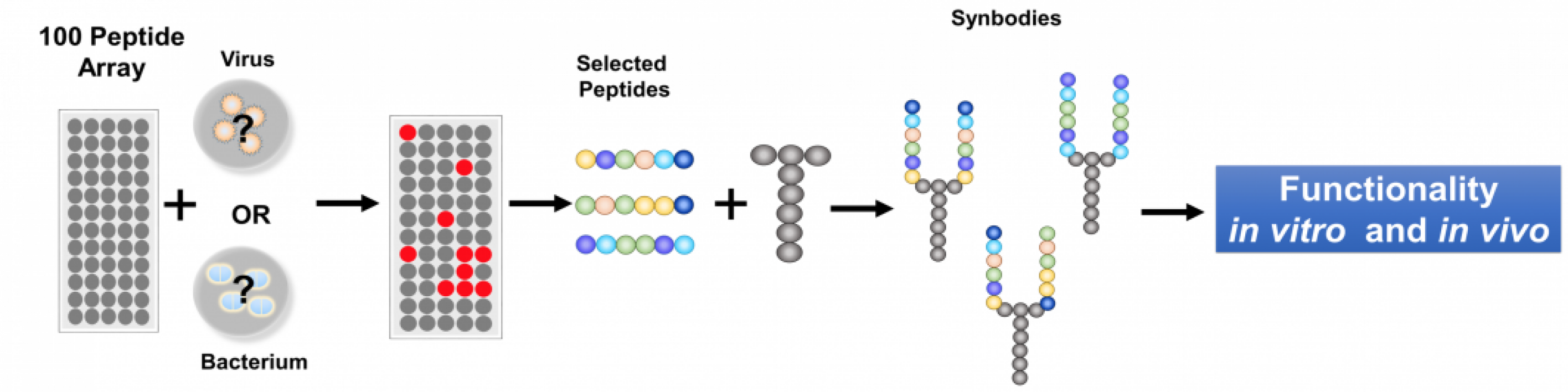
Synbodies are selected on peptide chips containing a premade set of 10,000 peptides placed in neat rows on a microscope glass slide, called a microarray.
To generate a synbody with antibiotic activity, a solution containing bacteria or viruses can be placed on the microarray.
“Our solution to save time was to pre-screen a large number of pathogens on the microarray and find 100 peptides that would be diverse enough so that any pathogen screened would bind to two or more peptides,” said Chris Diehnelt, an associate research professor in Johnston’s center who oversaw the lab experiments.
Stockpiling these 100 peptides in advance means that 1,000 or more doses of a therapeutic could be quickly produced after screening the best candidates that block a given pathogen. These candidates could then be produced in large amounts, purified and tested in mice for acute toxicity, completing the whole process in a week.
For their proof-of-concept, they screened a total of 21 different viruses and bacteria against their synbody arrays.
“We found that the majority of the peptides recognized one pathogen,” Diehnelt said.
In addition, they tested their system against two unknown pathogens that were not used in the study.
“The data showed that this array can potentially identify binding peptides for any given pathogen,” Diehnelt said.
A shot in the arm
“With this approach, dozens or even hundreds of synbodies can be produced in a day,” Johnston said.
The best candidates are quickly evaluated both for effectiveness and toxicity to humans and can be produced on a large scale.
Finally, their system was tested against two societal scourges and major worldwide health concerns: a potentially pandemic flu strain (H1N1 influenza) as a viral test and a bacterium that causes surgery-related infections, S. epidermidis.
“Our data indicate that a new virus or bacterium can be screened against the small peptide library to discover binding peptides that can be converted into neutralizing antiviral and antibacterial synbodies in a rapid manner,” Diehnelt said.
The next steps would be to prepare the final product for an IV delivery, and to scale up the system to produce enough product for a population-wide use — should the next Ebola or Zika or unanticipated outbreak occur.
“One key, unique feature of our synbody technology is that the same platform can produce synbodies with direct antibiotic or antiviral activity, and we can do it at a fraction of the potential cost as current, commercially produced therapeutic antibodies,” Johnston said.
This is good news: having a plan to save lives in place before the next inevitable outbreak.
And for Johnston, who has spun out companies based on other ASU technologies he has developed or co-developed, it could also mean big business. Global therapeutic monoclonal antibodies and cancer therapies are in high demand, with an estimated market size close to $100 billion for 2018.
This work was funded by a grant from the Defense Advanced Research Projects Agency (W911NF-10-0299) to Professor Stephen Albert Johnston under the 7-Day Biodefense Program.
Top photo: The H1N1 influenza virus. Image courtesy of the Centers for Disease Control and Prevention
More Science and technology

What does a spacecraft, a skeleton and an asteroid have in common? This ASU professor
NASA’s Lucy spacecraft will probe an asteroid as it flys by it on Sunday — one with a connection to the mission name.The asteroid is named Donaldjohanson, after Donald Johanson, who founded Arizona…

Hack like you 'meme' it
What do pepperoni pizza, cat memes and an online dojo have in common?It turns out, these are all essential elements of a great cybersecurity hacking competition.And experts at Arizona State…

ASU professor breeds new tomato variety, the 'Desert Dew'
In an era defined by climate volatility and resource scarcity, researchers are developing crops that can survive — and thrive — under pressure.One such innovation is the newly released tomato variety…