SMS Assistant Professor Guo decoding biological puzzles through single-cell proteomics

School of Molecular Sciences Assistant Professor Jia Guo.
Researchers across the country are looking for ways to successfully find the pieces to the biological puzzle of diseases like cancer and Alzheimer’s, as the demand for precise diagnosis and treatment options grow. Research in the field of single-cell proteomics is advancing quickly to try to meet those demands. The ability to study even just a few of the thousands of proteins in a single cell is helping scientists to analyze and decode some of those biological puzzles.
Researchers working in single-cell proteomics (SCP) are coming up with new ways to study and profile individual proteins in a single cell, keeping on pace with new technology as it evolves. The goal is to understand the complex spatial organizations, gene-expression regulations and interactions between the diverse cell types in multicellular organisms. Such understanding will provide valuable information on the detailed molecular-level mechanisms of diseases such as cancer.
One area of interest is the study of proteins in stem cells and the role they play in differentiation. If researchers are able to isolate and identify differentiation amongst these proteins in stem cells, areas like regenerative medicine could see benefits in tissue regeneration. Examples of tissue regeneration would be cartilage in knee repairs or tissue grafts in open heart surgery or skin grafts for burn victims.
At Arizona State University, Assistant Professor Jia Guo with the School of Molecular Sciences is one of several scientists looking at ways proteomics can help us understand disease development and specifically differentiation in stem cells. The Guo lab is developing methods to simultaneously detect and measure critical proteins present in individual cells of intact tissue using SCP.
Their method involves interrogating the tissue samples with an array of molecular detectors that contain antibody components that attach to specific and different target proteins. The detectors also contain a component that emits light with a color that is specific for the protein being detected. Under a microscope, the exact locations of the different proteins can be determined based on these different colors. After detection, the light-emitting component can be removed, via photocleavage, so that a different set of molecular detectors can then be added to the sample to probe for a different set of proteins. By using different rounds of this application/detection/cleavage method, a spatial map of a large number of different proteins can be obtained for individual cells in a tissue sample.
The importance of being able to perform this technique at the single-cell level is explained by Guo: “Unlike bulk-cell analysis, single-cell analysis can reveal cell heterogeneity, spatial organization, gene-expression regulation and interactions of diverse cell types."
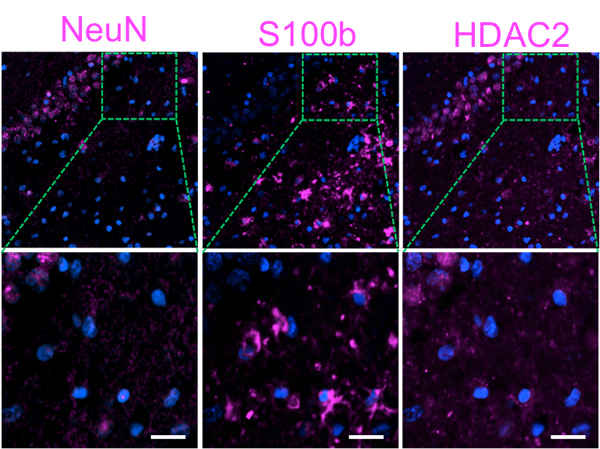
Three proteins are sequentially stained with cleavable Cy5 labeled antibodies in brain tissues. Nuclei are stained with DAPI (blue). Scale bars, 50 μm. (unpublished)
The Guo group has used this antibody-based SCP method to study the distribution of proteins in different kinds of brain tissue cells, including neuron and astrocytes. Based different spatial distributions of proteins, Guo and his group have found that the neurons in the human hippocampus can be partitioned into different kinds of clusters, and that different regions of the hippocampus are characterized by different kinds of neuron cluster.
To read more about Guo’s work with SCP, please find his interview with Technology Networks here: https://www.technologynetworks.com/cell-science/articles/through-the-looking-glass-of-single-cell-proteomics-310712
More Science and technology

SpaceHACK highlights student solutions to environmental challenges, digital divide
By Adrianna Nine About 250 students from around the world convened online and at Arizona State University on March 22 for the…

New AI for a new era of discovery
As the legend goes, in 1665, Sir Isaac Newton sat in his garden at Woolsthorpe Manor in England and looked on as a lone apple…

ASU receives 3 awards for research critical to national security
Three researchers in the Ira A. Fulton Schools of Engineering at Arizona State University have received grant awards under the …